| SLIT2 |
|---|
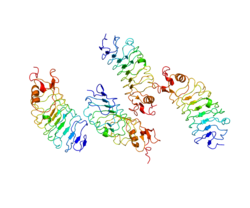 |
| Available structures |
|---|
| PDB | Ortholog search: PDBe RCSB |
|---|
| List of PDB id codes |
|---|
2V70, 2V9S, 2V9T, 2WFH |
|
|
| Identifiers |
|---|
| Aliases | SLIT2, SLIL3, Slit-2, slit guidance ligand 2 |
|---|
| External IDs | OMIM: 603746; MGI: 1315205; HomoloGene: 3516; GeneCards: SLIT2; OMA:SLIT2 - orthologs |
|---|
| Gene location (Human) |
|---|
 | | Chr. | Chromosome 4 (human)[1] |
|---|
| | Band | 4p15.31 | Start | 20,251,905 bp[1] |
|---|
| End | 20,620,561 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 5 (mouse)[2] |
|---|
| | Band | 5|5 B3 | Start | 48,140,480 bp[2] |
|---|
| End | 48,465,075 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - lower lobe of lung
- olfactory bulb
- vena cava
- trigeminal ganglion
- right lung
- spinal ganglia
- urethra
- pericardium
- parietal pleura
- middle temporal gyrus
|
| | Top expressed in | - floor plate
- ciliary body
- sciatic nerve
- vas deferens
- Epithelium of choroid plexus
- body of femur
- iris
- left lung lobe
- medullary collecting duct
- mammillary body
|
| | More reference expression data |
|
|---|
| BioGPS |  | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | - calcium ion binding
- heparin binding
- protein homodimerization activity
- GTPase inhibitor activity
- laminin-1 binding
- protein binding
- identical protein binding
- Roundabout binding
- proteoglycan binding
| | Cellular component | - cytoplasm
- membrane
- extracellular region
- cell surface
- extracellular exosome
- plasma membrane
- extracellular space
| | Biological process | - negative regulation of protein phosphorylation
- negative regulation of chemokine-mediated signaling pathway
- negative regulation of neutrophil chemotaxis
- negative regulation of cellular response to growth factor stimulus
- chemorepulsion involved in embryonic olfactory bulb interneuron precursor migration
- cell differentiation
- ureteric bud development
- negative chemotaxis
- negative regulation of actin filament polymerization
- negative regulation of small GTPase mediated signal transduction
- negative regulation of retinal ganglion cell axon guidance
- negative regulation of mononuclear cell migration
- corticospinal neuron axon guidance through spinal cord
- negative regulation of smooth muscle cell migration
- Roundabout signaling pathway
- positive regulation of axonogenesis
- nervous system development
- cell migration involved in sprouting angiogenesis
- negative regulation of endothelial cell migration
- axon guidance
- multicellular organism development
- chemotaxis
- negative regulation of smooth muscle cell chemotaxis
- cellular response to hormone stimulus
- negative regulation of cell migration
- negative regulation of monocyte chemotaxis
- negative regulation of cell growth
- response to cortisol
- apoptotic process involved in luteolysis
- positive regulation of apoptotic process
- induction of negative chemotaxis
- cellular response to heparin
- motor neuron axon guidance
- branching morphogenesis of an epithelial tube
- negative regulation of leukocyte chemotaxis
- chemorepulsion involved in postnatal olfactory bulb interneuron migration
- axon extension involved in axon guidance
- negative regulation of lamellipodium assembly
- negative regulation of vascular permeability
- retinal ganglion cell axon guidance
- negative regulation of GTPase activity
- aortic valve morphogenesis
- pulmonary valve morphogenesis
- ventricular septum morphogenesis
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | |
|---|
NM_001289135
NM_001289136
NM_004787 |
| |
|---|
NM_001291227
NM_001291228
NM_178804 |
|
|---|
| RefSeq (protein) | |
|---|
NP_001276064
NP_001276065
NP_004778 |
| |
|---|
NP_001278156
NP_001278157
NP_848919 |
|
|---|
| Location (UCSC) | Chr 4: 20.25 – 20.62 Mb | Chr 5: 48.14 – 48.47 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|

















